miR-ID® : accuracy through circularization
miR-ID® is a novel platform for detecting miRNA using a circularization-based RT-qPCR method. This technology allows SNP discrimination at any position in miRNAs and other small RNAs. miR-ID® is highly sensitive and uses affordable single-dye detection. The technology works well with all sample sources, including total RNA, cell lysates, and tissue lysates.
miR-Direct® : miR-ID® - powered miRNA quantification directly from serum or plasma
SomaGenics' miR-Direct® method overcomes commonly encountered problems of inconsistent RNA recovery, PCR inhibition by sample stabilizers, and weak signals caused by low RNA abundance in biofluids. miR-Direct®- based miRNA capture is followed by miR-ID® quantification. Accurate quantification of microRNAs (miRNAs) from biofluids is crucial to realizing their potential as potential biomarkers.
miR-ID® Technology overview
miR-ID® is highly sensitive and specific
For example, the closely related miRNAs 34a, b, and c are well discriminated by miR-ID®. To determine the relative Ct values listed in the table, the difference in Ct values for the intended target and their related isoforms was converted into a linear percentage, with the fully matched reaction normalized to 100%. All non-specific detection is well below 1% for these assays.
The dilution curves indicate that all three assays have a dynamic detection range of 8 logs.
miR-ID® detects end-modified small RNAs
miR-ID® quantitatively discriminates between small RNAs with an identical sequence but differing terminal modifications. This is particularly useful in plant miRNAs, where discrimination between 2-O-methyl modified and unmodified miRNA is of considerable interest.
miR-ID® can quantitatively detect 2' O-methyl modifications at the 3’-terminus of miRNAs. By selecting enzymes that circularize either both types of miRNAs (with 2’-OMe or 2’-OH forms on the 3’termini), shown in green, or 2’-OH forms only (shown in red), these two end-modifications can be distinguished. In this example, synthetic let-7b miRNAs with the two respective 3’ termini were assayed and quantification with the enzyme circularizing both forms were set to 100%
miR-ID® technology schematic
Circularization of linear miRNAs is followed by rolling circle amplification during the reverse transcription. The circularization step allows placement of RT primers at any location of each miRNA to optimize discriminating single nucleotide-polymorphisms (SNPs) at any location of the miRNAs.
Rolling-circle reverse transcription generates multimeric cDNAs. This method allows that primers that are entirely complementary to the miRNA sequence can be used for the real-time PCR, therefore increasing PCR specificity.
Detection of the real-time PCR signal is through SYBR chemistry.
miR-ID® workflow
All steps up to quantitative PCR are performed in multiplex in a single tube.
miR-Direct® Technology overview
miR-Direct® yields optimal results for plasma and serum
Plasma and serum from the same collection were analyzed by the indicated preparatory and qPCR methods. miR-Direct® samples accommodated a 200 ul input of biofluids, whereas the recommended 8 ul input was used for the column preparations. miR-Direct® is not inhibited by heparin and miRNA quantifications are similar from plasma and serum.
miR-Direct® is more sensitive than a LNA-based method
miR-Direct® was directly compared to a LNA-based method using 200ul plasma input of the same donors and analysis of two different miRNAs.
This analysis was performed by unaffiliated researchers.
miR-Direct® reliably detects miRNA in urine samples
miR-Direct® enables miRNA detection with higher sensitivity than the competitor kit. All Ct values over 37 are considered background level (not detectable).
miR-Direct® technology schematic
miRNA is directly captured from variable input volumes (25 to 400 µl), eliminating solvent extraction or column purification. This process removes inhibitors of enzymatic reactions, including heparin.
Up to the PCR step in the miR-ID® procedure, the entire process occurs in a single tube.
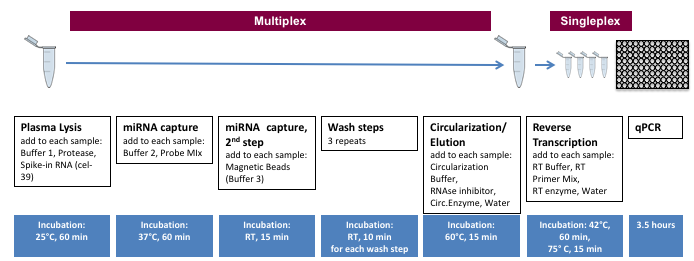
miR-Direct® workflow
All steps up to quantitative PCR are performed in multiplex in a single tube. Note that this schematic includes the miR-ID® steps.
Legal and Patent Information
These products are covered by one or more patents, trademarks and/or copyrights owned or controlled by SomaGenics, Inc. No right under any other patent claims to methods, apparatus or reagents is conveyed expressly, by implication, or by estoppel. This product is intended for Research Use Only and may not be transferred to third parties, resold, modified for resale or used to manufacture commercial products without prior written approval of SomaGenics, Inc. For more information about commercial rights, please contact SomaGenics.